Biologist Stephen Friend is president and co-founder of the nonprofit research organization Sage Bionetworks in Seattle, and co-director of the Resilience Project, which starts in September He is searching for exceptional people who carry genes for serious childhood disease but have never gotten sick—are you one of them?
Why do you want to find people who have genes for serious diseases?
We aim to find relatively healthy adults who have somehow escaped having the classical symptoms, because although they carry the faulty gene that would normally mean they get the disease, they also have something else protecting them—possibly another mutation. We know people who have made it to adulthood without any symptoms are going to be very rare. That’s why we are referring to them as unexpected heroes.
How do you know these people are out there?
My partner in this project Eric Schadt, a genomics professor at Mount Sinai Hospital in New York, has seen examples. One was a woman in her 50s with a mutation in the gene that causes cystic fibrosis but who has never had anything more than mild respiratory problems. And there was a 45-year-old man who never had any symptoms but learned by chance that he had the usually fatal neurological condition Louis-Bar syndrome, which is caused by a faulty gene. Both had an inherited childhood disease but no symptoms. That is exactly what we are looking for.
How will you find these resilient people?
We are asking people to volunteer their DNA in a systematic worldwide search, looking at the vast majority of inherited childhood diseases—from cystic fibrosis to Rett syndrome.
Why is finding them so important?
We know that those who avoided getting sick when young may harbor protective genes. The assumption is that finding these protective factors is a very direct path to developing new therapies.
That’s because almost all of the genetic alterations that cause disease are due to a loss of function, say in a protein that the faulty gene codes for. Equally, a loss of function caused by the second mutations our unexpected heroes carry is probably what’s providing protection. The vast majority of drugs we have use small molecules, which are extremely poor at restoring function, but very good at disrupting it. So if you found a dozen of these protective factors, there is a high chance you could build drugs to mimic most of them by recreating the loss of function.
Who can take part in the project?
Anyone who is over 30 and was relatively healthy in childhood—who did not go to a doctor with a severe illness—can volunteer. None of the hundreds of inherited diseases we are considering would be a minor bother.
How can people contribute their genetic information?
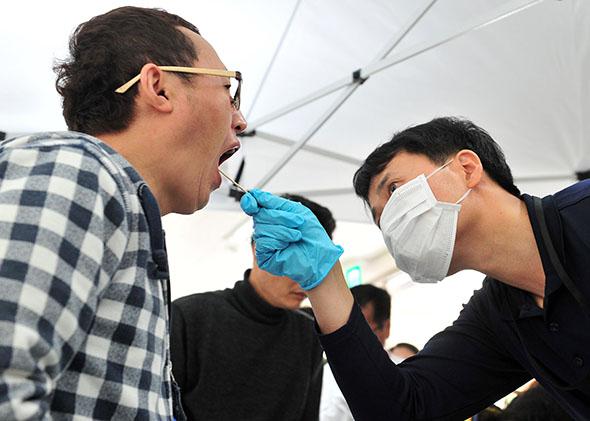
You sign up and give consent on our website resilienceproject.me—you can register interest in taking part now. We will send out a cotton swab that you scrape inside your cheek to pick up enough cells to allow your DNA to be analyzed. You mail it in, and we test for the 125 childhood diseases we’re focused on—those that are very severe, where you are highly likely to suffer the full illness if you have the mutated gene. After we test your sample, you are most likely to get a card saying that we don’t see any evidence that you are an unexpected hero. Or it may say we think it is worth doing further analysis to validate something we found.
We hope to find a million people to take part, and among them, the handful of people who are likely to fit the unexpected hero criteria.
What will you do when you find this handful of people?
To see if we can find their resilience factors, we would want to look deeply at their whole genome and other aspects of their biology that might provide clues, such as their RNA, proteins, and metabolism.
It is like trying to find a needle in a haystack. And to make it more stark, we are not expecting to have hundreds of such individuals for any of the illnesses we are targeting, which might have made it easer to find these protective factors. Instead, we will have isolated individuals with a variation that has occurred somewhere in their massive genome. So it won’t be easy, but methods have emerged in the last five years to allow us to attempt it.
What are those methods?
One is called network biology filtering. Instead of treating every piece in the genomic haystack as one of a million to look through, we can focus more narrowly in the network connected to a faulty gene.
The other approach involves using gene-editing techniques. We can create the primary mutation, and other possible buffering mutations one or two at a time, in a cell line in the lab, then work through them to find what offers protection.
Has the idea of finding treatments by studying those who don’t get sick been tried before?
Yes, but not in the systematic genetic way that we are doing, which was too costly until now. There are two earlier efforts that encouraged us. One is a genetic approach: For most inherited diseases, scientists and clinicians have studied an extended family with a particular mutation, looking for individuals who carried the faulty gene but had fewer symptoms. The other approach uses observable traits. For example, Helen Hobbs at the Howard Hughes Medical Institute in Maryland sought patients with high lipid levels who did not get heart disease. She discovered mutations in a gene that result in reduced levels of “bad” LDL cholesterol. That is now being built as a therapy for heart disease.
How have you prepared for the project?
Before we launched, we thought it would be a good idea to look at existing samples in anonymized databanks, such as 23andMe, to get an idea of whether there would be any unexpected heroes. About half of the disease-linked genes we were interested in analyzing were represented. We screened half a million samples and found dozens of unexpected heroes—people that appear to have, for example, the genetic fault that causes cystic fibrosis, but are generally healthy.
You are hoping to have volunteers from around the globe. Why is that so important?
Resilience in the populations we have looked at so far is very infrequent, occurring in one in 35,000 individuals. So how might you increase the chances of finding these unexpected heroes? We think there are several ways. One is to go to regions of the world where first cousins marry first cousins; that brings up the frequency of alterations. In the Middle East, for example, if we enroll 100,000 people, there is a chance that we will find more unexpected heroes than in the U.S. We are also hoping that there are environmental niche populations, whether in the Amazon or in the Arctic, that may carry protective factors genetically or as a result of environmental influences that will again be of benefit for finding resilience.
Is it possible to pinpoint environmental factors that might help protect against these diseases?
That is much harder. Most environmental studies rely on large numbers. We are not going to have those numbers. The reality is that for the nongenetic factors, entirely separate studies would need to be done afterwards to try and find hamlets or villages where there is a disproportionate effect, where you might be able to identify an environmental cause.
When might the project yield new treatments?
Finding unexpected heroes is likely to take a few years, but I don’t think it should take more than a year to decipher their data. Then, to turn an intriguing target that we hopefully find into a possible treatment, I think it would be another five years to get to clinical trials. It pretty much adds up to a decade.
But to get there, most of all we need individuals to step up and volunteer. It just takes a swab of DNA and a willingness to wonder, “What’s inside me?”
This article originally appeared in New Scientist.